Bim package: Difference between revisions
No edit summary |
No edit summary |
||
| Line 3: | Line 3: | ||
The data for this example can be found in the doc directory inside the | The data for this example can be found in the doc directory inside the | ||
bim installation directory. | bim installation directory. | ||
We want to solve the equation | |||
<math> -\mathrm{div} ( \alpha \gamma ( \eta \nabla u - \beta u ) )+ \delta \zeta u = f g </math> | |||
<b> Create the mesh and precompute the mesh properties </b> | <b> Create the mesh and precompute the mesh properties </b> | ||
| Line 38: | Line 43: | ||
<b> Set the coefficients for the problem:</b> | <b> Set the coefficients for the problem:</b> | ||
Revision as of 16:18, 19 July 2012
This is a short example on how to use bim to solve a DAR problem.
The data for this example can be found in the doc directory inside the
bim installation directory.
We want to solve the equation
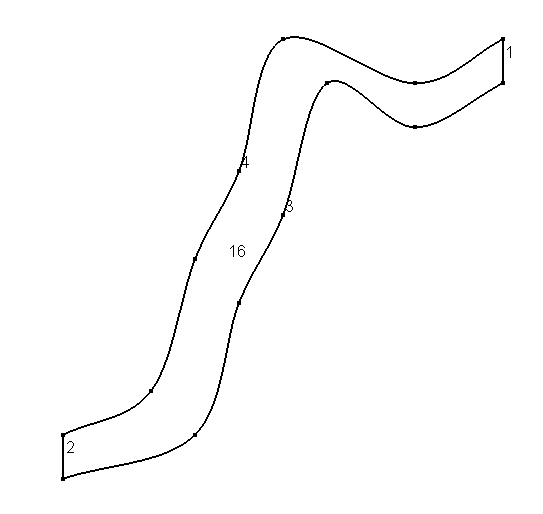
Create the mesh and precompute the mesh properties
The geometry of the domain was created using gmsh and is stored in the file fiume.geo created with gmsh
[mesh] = msh2m_gmsh("fiume","scale",1,"clscale",.1);
[mesh] = bim2c_mesh_properties(mesh);
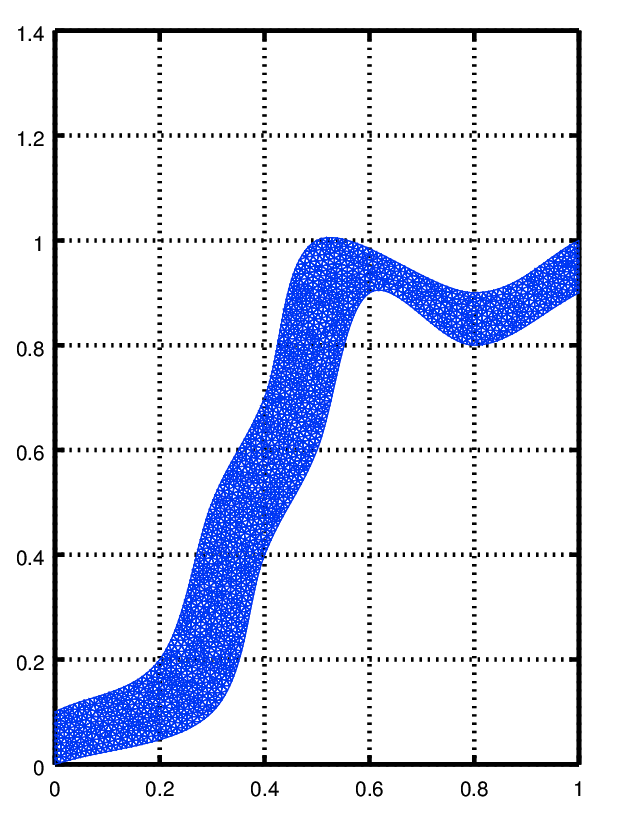
to see the mesh you can use functions from the [fpl] package
pdemesh (mesh.p, mesh.e, mesh.t) view (2)
Construct an initial guess
For a linear problem only the values at boundary nodes are actually relevant
xu = mesh.p(1,:).'; yu = mesh.p(2,:).'; nelems = columns(mesh.t); nnodes = columns(mesh.p); uin = 3*xu;
Set the coefficients for the problem:
epsilon = .1; alfa = ones(nelems,1); gamma = ones(nnodes,1); eta = epsilon*ones(nnodes,1); beta = xu+yu; delta = ones(nelems,1); zeta = ones(nnodes,1); f = ones(nelems,1); g = ones(nnodes,1);
Construct the discretized operators
AdvDiff = bim2a_advection_diffusion(mesh,alfa,gamma,eta,beta); Mass = bim2a_reaction(mesh,delta,zeta); b = bim2a_rhs(mesh,f,g); A = AdvDiff + Mass;
To Apply Boundary Conditions, partition LHS and RHS
The tags of the sides are assigned by gmsh
Dlist = bim2c_unknowns_on_side(mesh, [8 18]); ## DIRICHLET NODES LIST Nlist = bim2c_unknowns_on_side(mesh, [23 24]); ## NEUMANN NODES LIST Nlist = setdiff(Nlist,Dlist); Fn = zeros(length(Nlist),1); ## PRESCRIBED NEUMANN FLUXES Ilist = setdiff(1:length(uin),union(Dlist,Nlist)); ## INTERNAL NODES LIST
Add = A(Dlist,Dlist); Adn = A(Dlist,Nlist); ## shoud be all zeros hopefully!! Adi = A(Dlist,Ilist); And = A(Nlist,Dlist); ## shoud be all zeros hopefully!! Ann = A(Nlist,Nlist); Ani = A(Nlist,Ilist); Aid = A(Ilist,Dlist); Ain = A(Ilist,Nlist); Aii = A(Ilist,Ilist); bd = b(Dlist); bn = b(Nlist); bi = b(Ilist); ud = uin(Dlist); un = uin(Nlist); ui = uin(Ilist);
Solve for the displacements
temp = [Ann Ani ; Ain Aii ] \ [ Fn+bn-And*ud ; bi-Aid*ud]; un = temp(1:length(un)); ui = temp(length(un)+1:end); u(Dlist) = ud; u(Ilist) = ui; u(Nlist) = un;
Compute the fluxes through Dirichlet sides
Fd = Add * ud + Adi * ui + Adn*un - bd;
Compute the gradient of the solution
[gx, gy] = bim2c_pde_gradient(mesh,u);
Compute the internal Advection-Diffusion flux
[jxglob,jyglob] = bim2c_global_flux(mesh,u,alfa,gamma,eta,beta);
Save data for later visualization
fpl_dx_write_field("dxdata",mesh,[gx; gy]',"Gradient",1,2,1);
fpl_vtk_write_field ("vtkdata", mesh, {}, {[gx; gy]', "Gradient"}, 1);